Our mission
Cancer is a global health burden. In-vitro pre-clinical models play a key role in fighting this burden by encompassing all the activities prior to clinical trials, from tumor microenvironment reconstruction to drug candidate selection. However, the frequent failure of promising pre-clinical drug candidates highlights two major drawbacks of these models: (i) the difficult reproduction of the dynamic cancer structure related to numerous physical cues; (ii) their experimental nature that suffers from high costs, long times, and limited understanding. Consequently, the relationship between dynamic physical cues, cell behavior, and drug efficacy is still unknown.
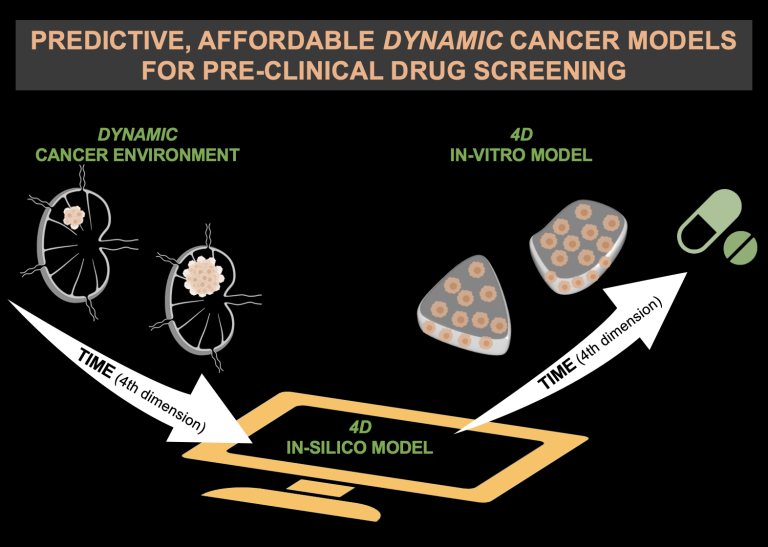
CoDe4Bio tackles such a huge knowledge deficiency: it proposes a radical methodology shift to a computational approach to harness programmable materials, able to change properties on demand and realize dynamic 4D biofabricated models whose stimuli-triggered evolution over time (4th dimension) induces targeted physical cues on cancer cells.
The research team will leverage the Principal Investigator extensive experience with smart materials and structures to address the challenges of this multidisciplinary project. Specifically, the team will develop a computational design framework for 4D biofabrication that combines new data-, geometry-, and model-based methods with additive manufacturing and in-vitro observations. This framework will allow the researchers to develop customized stimuli-responsive materials and engineer a new generation of 4D constructs with programmable mechano-structural properties acting as mechanical regulators.
The researchers will also assess the constructs in-vitro on chronic lymphocytic leukemia to achieve a deep understanding on how complex physical cues within lymph nodes and bone marrow affect this incurable cancer in relation to chemoimmuno and targeted therapies.
CoDe4Bio will push the frontiers of solid and computational mechanics to unveil unconventional routes for pre-clinical drug screening and lay the foundation for effective dynamic cancer models.
